The problem
At Finch Therapeutics, which developed microbiome therapeutics, the scientific platform needed to select microbes derived from human stool to be included in a Live Biotherapeutic Product for the treatment of Ulcerative Colitis. After tens of thousands of selections and passages, the microbiologists ended up with freezers of potential candidates–which half-dozen microbial isolates should they select?
The platform
Right Bionic helped Finch construct their digital isolate library. We partnered with their computational biologists to deploy a pipeline that could rank the similarity of any two isolates based on genetic barcodes. Whenever the bench scientists handled new microbes, the pipeline was run automatically and the classification results were stored in a data warehouse, with unique IDs assigned. The bench scientists could see their clusters of microbes via a web UI, and the data scientists could pull the results into their notebooks via an API that Right Bionic developed. Both bench and data scientists then had a shared nomenclature for selecting which isolates to continue work on.
This resulted in the first internally developed drug candidate nomination.
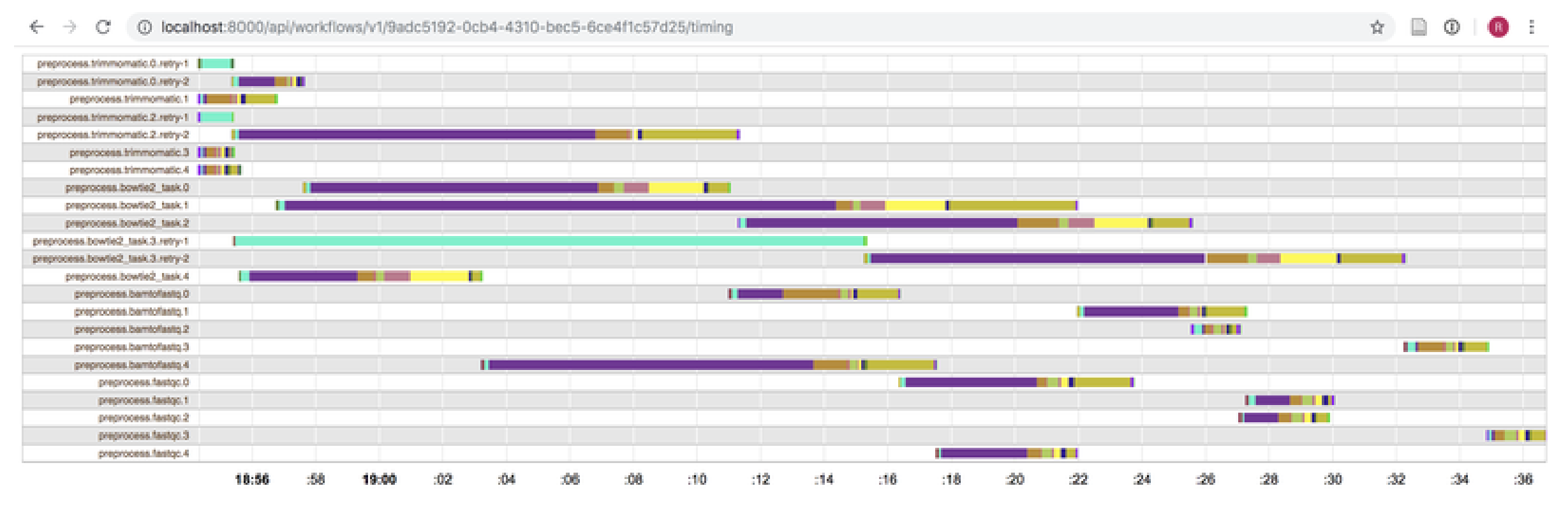
Recipt, in technical terms
Ingredients:
- Benchling Bioregistry
- Google Cloud Run (i.e. Knative)
- Cromwell Engine/Google Life Science API
- Airflow (ETL orchestrator)
- Identity Aware Proxy (for securing web apps)
- Cloud Bucket
- Big Query
Step 1, whenever the microbiologists inoculate a new culture, have the microbiologists register it in Benchling as a new child entity. Sequence its 16S gene (which can be used like a barcode for microbial identity). In this case, we sent it out to a CRO, who shared the data back with us via their website.
Step 2, every night (scheduled by Airflow orchestration software), run a job that scrapes the CRO website and downloads the data into S3.
Step 3, After the data have been ingested, tag the files with the Benchling entity ID to establish its provenance. We used a custom software called the Raw Data API (really just a metadata database that pointed to files in S3).
Step 4, With every new Sanger sequencing file, spin up a Cloud Run job that QCs and aligns the sequence to that of all the microbes' ancestors. Store the results in a database, made accessible by an API.
Step 5, Spin up a simple flask web application that displays the data from Benchling + Raw Data + Genetic similarity. Microbiologists can put in a Benchling entity ID and see whether it is the same or different than its ancestor (if it's different ... that's a different discussion)
Step 6, Give your data scientists a simple python package that lets them consume the data APIs including Benchling via their notebooks. This way everyone has the same data, delivered in the way they want it.
Step 7, Assign IDs to the continuous microbes. In the results database, cluster all similar microbes according to Algorithm A. Call the microbes something like AlgoA-1, to differentiate it from any other type of nomenclature.
Contact
Ryan Bellmore
[email protected]
rightbionic.com
Let's talk about your data.
